Med-Chem Optimization of Paxlovid
Tagged:COVID
/
PharmaAndBiotech
Previously, we wrote about how hard it currently is to find paxlovid, given intense demand, supply chain frustrations, and the complex synthesis pathway. Today let’s have a look at how exactly one goes about discovering such a thing, with an improbable number of steps in its chemistry. (Hint: an awful lot of luck/serendipity was involved!)
Science… and serendipity
Of course, an enormous amount of effort in drug discovery goes into steps that occur before chemistry (long before, as in 5 or so years of effort). A slimmed-down, overly simplified version of the research pipeline that might feed into the development pipeline is something like:
- Identify diseases that have theories for their mechanisms that look reasonably tractable with either a small molecule drug, an antibody, an antibody/drug conjugate, a protein degrader, immune system attack, or whatever other trick of the day you happen to have in your back pocket.
- Prioritize the diseases based on unmet medical need.
- Identify targets, usually proteins, that when inhibited will modify the disease. (Sometimes the targets are not proteins, and very rarely you enhance rather than inhibit protein activity).
- Target validation in which it’s proven rock-solid that modifying the target modifies the disease, in cell cultures, organoids, and possibly a mouse or rat model.
- Pick a mechanism of attack:
- CRISPR/siRNA/shRNA or the like to prevent the protein being made,
- a small molecule to glom onto a pocket in the protein and gum up its function,
- an antibody to bind to the protein (if on the cell surface, possibly to signal the immune system to kill that cell),
- an antibody/drug conjugate (e.g., deliver a toxin exactly and only to cells expressing your protein target on the cell surface)
- immune system modulators (traffic killer T cells into an otherwise “cold” tumor, bispecific antibodies to bind to a cancer cell antigen and a killer T cell so the immune system learns to kill things like that, and so on),
- a protein degrader (binds to the protein in question and a ubiquitin ligase, so the protein gets ubiquitinated and thus flagged for destruction in the proteasome),
- synergistic combinations of the above, in which 2 treatments produce more effect than their separate sums (including combination with other things like radiation therapy),
- … and maybe a dozen other mechanisms
- Then, and only then, you call in the miracle workers in the form of your friendly local med-chem folks. I used to really like these guys. They were smart, funny, and regularly delivered miracles in the form of nanomolar- or even femtomolar-potency molecules that they could synthesize regularly and reliably.
- Only at this point, after maybe 7 or 8 years of work in most cases, do you even begin to talk to the clinical people about how maybe they might want to take an interest in a clinical trial? (And yes, weird stuff can happen: I’ve seen the 7 or 8 years of work of maybe 50-100 scientists get dropped at this point, because the clinical folk didn’t think it was fashionable.)
Pfizer, when looking for COVID-19 therapeutics, got extremely lucky: previous work on SARS-CoV1 in China in the 2003 had already finished steps 1-5. They had a candidate molecule, PF-00835231, already at least partially optimized in med-chem, ready to move into clinical trials. But then SARS burned itself out, to the project got shelved. I’m sure there was some nervous side-eye when MERS broke out in the Middle East with its high mortality rate, but thankfully that burned out too.
Still, they shelved the project, didn’t throw it out. Yes there probably were knoweldge preserving systems, but more importantly there was institutional knowledge, in the form of people who still knew all this (and could probably show you the battle scars). So when SARS-CoV2 broke out into COVID-19 in late 2019, they remembered.
They knew that this coronavirus family had the giant gene ORF1ab. It encodes a “polyprotein”, i.e., a long string comprising several distinct viral proteins. The virus depends on a protease, called 3CLpro or Mpro, to cleave this apart into the functional proteins. If you gum up 3CLPro, then the polyprotein is not cleaved and the virus is stopped from reproducing. More importantly, there’s nothing much like 3CLpro in human, so you’re preferentially attacking an essential target in the virus, but not a human target. This makes 3CLpro a very attractive therapeutic target because it has a huge dose window: you can slam the virus really hard before you start having much effect on human cells. They already had molecules that did this for the original SARS virus from 20 years ago.
And that, boys and girls, is a helluva good starting point! On 2020-Mar-13, Pfizer med chemist Dafydd Owen was sent home from the lab in Cambridge, Massachusetts to work from home in the pandemic. He as told basically to “take a weekend” to think through how he’d put together a team to make a COVID-19 therapeutic. Owen, knowing where the bodies were buried, of course wisely chose the previous 3CLpro SARS inhibitor as a starting point. That is how you get t a human clinical trial in less than 12 months: start with a mostly-ready drug for a related disease, crash-priority funding and more or less unlimited resources, and a dedicated team of smart (and slightly desperate) people.
 And that’s today’s story, brought to us by masterful med-chem blogger Derek Lowe at In
the Pipeline [1], starting from an article in Chemical
& Engineering News by Bethany Halford. [2]
And that’s today’s story, brought to us by masterful med-chem blogger Derek Lowe at In
the Pipeline [1], starting from an article in Chemical
& Engineering News by Bethany Halford. [2]
They document the gnarly med-chem optimization work that led to nirmatrelvir, one of the 2 molecules that comprise paxlovid. (The other is ritonavir. It’s also a protease inhibitor used in HIV and hepatitis C. But more importantly, it inhibits the liver enzyme CYP3A that degrades many drugs for elimination from the body. On the one hand, this increases the lifetime of nirmatrelvir in the body and makes it stronger; on the other hand, it interacts with nearly every drug you’re like to take so physicians have to examine your prescriptions very carefully to modify them for paxlovid therapy.)
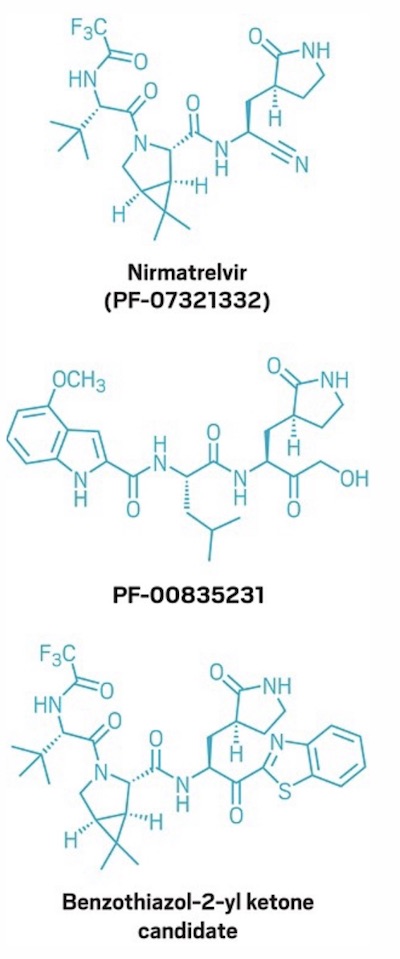 C&E News reported this set of structures. The middle one, PF-00835231, is the lead
candidate for the original SARS project back in 2003. It’s similar to a peptide that
binds to 3CLpro in nature. It binds to the 3CLpro ortholog in SARS-CoV1 well enough, but
they wanted to do better. The bottom molecule is one of the intermediate candidates
— these guys build anywhere from dozens to hundreds of these, searching for
molecules that work better and are easier to synthesize — and the top molecule is
the current nirmatrelvir structure.
C&E News reported this set of structures. The middle one, PF-00835231, is the lead
candidate for the original SARS project back in 2003. It’s similar to a peptide that
binds to 3CLpro in nature. It binds to the 3CLpro ortholog in SARS-CoV1 well enough, but
they wanted to do better. The bottom molecule is one of the intermediate candidates
— these guys build anywhere from dozens to hundreds of these, searching for
molecules that work better and are easier to synthesize — and the top molecule is
the current nirmatrelvir structure.
But, of course, there were problems:
- PF-00835231 is rich in hydrogen bonds, so those 5 H bonds will trap it in the GI tract,
preventing it from being orally available. It has to be given by IV infusion, which is
ok in good times, but out of the question when hospitals are near collapse.
- But some of those bonds are useful, e.g., the $\alpha$-hydroxymethyl ketone binds to a cysteine in 3CLpro. So you need to replace it with an equivalent cysteine binder. One choice led to the benzothiazol-2-yl ketone in the bottom structure, and the other was a nitrile. Only after much testing was the nitrile picked.
- There’s another H bond at the leucine moiety. Long story short: replaced with a cyclic amino acid that looks a bit like leucine to be attractive to 3CLpro. They used a fused cyclopropyl ring with a couple methyl groups, based on precedent in another antiviral drug (Schering-Plough’s hep C drug boceprevir).
- But… gaack… if you do all that, you mess up other stuff. In this case, you lose contact with a glycine in the 3CLpro binding pocket. So they started messing with the indole moiety to fix this, with one thing and another: methyl sulfonamide, acetamide, they even tried Raoul Mitgong, but he didn’t help much… before settling on that incendiary-looking triflouroacetamide in the upper left. Nobody’s first choice, but it works.
- Finally they had to choose between the nitrile and the benzothiazol-2-yl ketone. The nitrile won because it’s more soluble (sine qua non of orally available meds), it’s chirally more stable (doesn’t fold up the wrong way), and the scale-up chemistry to manufacture it by the ton was more feasible. (As a scale-up chemist once said, if not exactly to me, at least in my hearing: “Sure, you lab weenies can make a few milligrams or even grams of the stuff. But I gotta make tons of it. Throw me a bone, here!”)
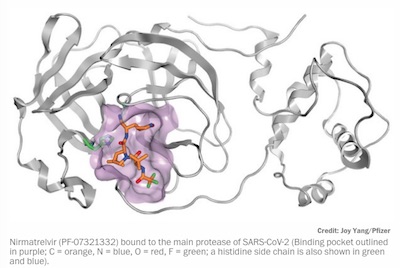 On 2020-Jul-22, they first made PF-07321332, now known as nirmaltrelvir, as one of about
20 compounds to be tested. The rat PK studies came back 2020-Sep-01, and this molecule
was blessed. Here, in another illustration from Joy Yang of PFizer via C&E News, it’s
shown in its binding pocket in 3CLpro:
On 2020-Jul-22, they first made PF-07321332, now known as nirmaltrelvir, as one of about
20 compounds to be tested. The rat PK studies came back 2020-Sep-01, and this molecule
was blessed. Here, in another illustration from Joy Yang of PFizer via C&E News, it’s
shown in its binding pocket in 3CLpro:
- The gray arrows and curves are the 3CLpro protein structure.
- The binding pocket is outlined in purple, to give you an idea where the electron density is.
- For nirmatrelvir, carbons are orange, nitrogens are blue, oxygens are red, and fluorines are green. The histidine side chain hanging off the back is also shown in green & blue.
This is what you want to see: an nice, deep pocket in the target protein, essential for its function, tightly gummed up with a small molecule of your crafting which prevents the protein from functioning.
At that point, the “lab weenies” hove to and made 1.4 kg of nirmatrelvir for the clinical trial. There are some very gnarly starting materials, like the bicyclic structure and the lactam, but at least there’s a plausible supply chain for those. We hope.
It was fast at 12 months from standing start to clinical trial. But it started from a very advanced point, based on previous work. And it was the work of literally 1000s of scientists working flat-out with no breaks, and with an apparently nearly unlimited budget. But here we are: an orally available protease inhibitor combination drug, which has about 89% efficacy in preventing hospitalization.
That… is good work. I’ve never seen anything that even comes close to this. We can’t repeat the lucky starting point, but we can appreciate that you never let go of prior research.
Activity vs the SARS-CoV2 Omicron variant
 The other question everybody wants to know about: will this work against Omicron, as well?
The answer should be yes, because while Omicron has many mutations, very few are in
3CLpro, the target of nirmatrelvir. Along those lines came a press release yesterday from
Pfizer [3]:
The other question everybody wants to know about: will this work against Omicron, as well?
The answer should be yes, because while Omicron has many mutations, very few are in
3CLpro, the target of nirmatrelvir. Along those lines came a press release yesterday from
Pfizer [3]:
NEW YORK, January 18, 2022 — Pfizer Inc. (NYSE: PFE) today shared results from multiple studies demonstrating that the in vitro efficacy of nirmatrelvir, the active main protease (Mpro) inhibitor of PAXLOVID™ (nirmatrelvir [PF-07321332] tablets and ritonavir tablets), is maintained against the SARS-CoV-2 variant Omicron. Taken together, these in vitro studies suggest that PAXLOVID has the potential to maintain plasma concentrations many-fold times higher than the amount required to prevent Omicron from replicating in cells.
So it looks like we can hope for some pan-coronavirus activity, given conserved structure of viral protease 3CLpro (Mpro).
The press release goes on for some time, as these things tend to do. Much of it is
gluteal armor, to prevent them from getting bitten in the behind by lawyers now that this
is a medication being given to actual people. But stripped of that, there were a few
other interesting things about in vitro studies of nirmatrelvir across SARS-CoV2 variants:
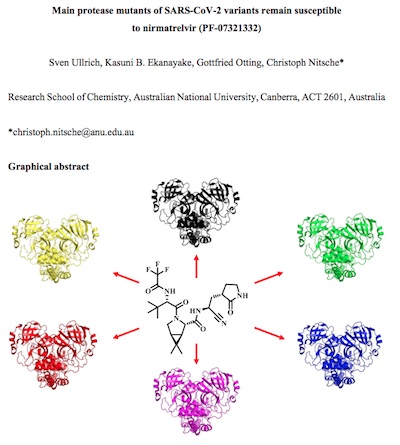
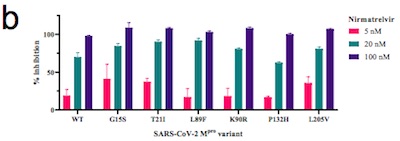
- It looks like the binding affinity of nirmatrelvir for 3CLpro is very good at $K_i \sim 1\mbox{nm}$, for both the original SARS-CoV2 and the Omicron variant. They alleged a preprint available on bioRχiv, but didn’t cite anything in particular, in the frustrating manner customary in press releases. However, a bit of spelunking leads me to believe it’s this one by Ullrich, et al. [4]. They looked at the activity of nirmatrelvir against 3CLpro in 6 different SARS-CoV2 lineages, including Omicron, and found good activity everywhere. It looks like, from their Figure 2b, that there’s > 50% inhibition at doses below 20nM for all variants examined. That’s good news.
- In a second in vitro study, viral loads were measured vs dose and an EC50
(concentration at which half the effect was achieved) calculated. They looked at this
dose response in Omicron and other variants, getting around 16nM vs 38nM in original
strain, i.e., better response in Omicron. Again they claim to have submitted to
bioRχiv, but this being a press release they couldn’t be bothered to give a proper
citation. Spelunking on my part was less successful, but this appears to be a related
result by Bojkova, et al. [5]
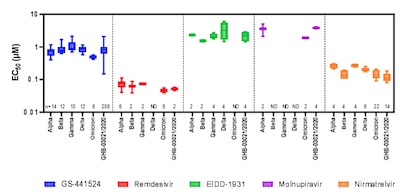
- A third in vitro study compared nirmatrelvir and “other authorized/approved COVID-19 therapeutics”. (This being a press release, they obstinately refuse to name the others.) They got IC50s (concentration at which 50% of infections were stopped) of 38 - 207nM for nirmatrelvir against Omicron vs 22 - 225nM against the original strain, i.e., entirely comparable. Again, this being a press release, they claim to have submitted to bioRχiv but give no citation. However, they do cite other work by Vangeel et al. [6], showing their “findings are consistent.” (Why in the world would you fail to cite your own research, but instead cite someone else’s as “consistent” with your own?!) However, the Vangeel paper compares remdesivir, molnupiravir, and nirmatrelvir in various SARS-CoV2 variants and indeed comes to about that conclusion. Shown here is their Figure 1, with IC50’s of nirmatrelvir vs viral variants on the far right.
The Weekend Conclusion
It looks like the rapid development of paxlovid was due to a combination of luck (previous work on a 3CLpro inhibitor for SARS in 2003) and hard work (Owen and his team at Pfizer). Another piece of luck is the broad activity against SARS coronavirus variants, which is very hopeful indeed.
Some of this was serendipity, with the previous 3CLpro inhibitor being remembered at the right time by the right people. As I’ve been known to say, somewhat irritatingly:
You can’t plan serendipity – give chance a chance!
… and here serendipity came through for us. There’s nothing so useful as a huge knowledge base of what you’ve tried in the past, to inform what you should try next.
The religious side of me wants to be grateful for whatever divine influence put these tools in our hands at precisely the right moment. The more pragmatic side of me just wants all of us to be thankful for the fantastic series of fortunate coincidences:
- mRNA vaccine research, right on the cusp of feasibility when COVID-19 hit
- Protease inhibitor research on coronaviruses, just sitting there ready to be repurposed
- Previous work on SARS1 and MERS, complete with starting molecules for SARS, ready for med-chem optimization
- Pretty good efficacy against Omicron of mRNA vaccines and paxlovid (both vs hospitalization, not vs even minor infection)
- Internet infrastructure: video calls, remote work (at least for some), food delivery, telemedicine … imagine how bad isolation would have been in the 1970s without any of that!
We have a lot to regret (millions dead, a worldwide descent into superstitious vaccine defiance coupled with revenant right-wing fascism), but we also have a lot for which we can be very grateful.
I prefer the latter.
Notes & References
1: D Lowe, “Discovering Paxlovid”, In the Pipeline blog at Science Translational Medicine, 2022-Jan-18. ↩
2: B Halford, “How Pfizer scientists transformed an old drug lead into a COVID-19 antiviral”, Chemical & Engineering News, 100:3, 2022-Jan-14. ↩
3: Pfizer Media Relations, “Pfizer Shares In Vitro Efficacy of Novel COVID-19 Oral Treatment Against Omicron Variant”, Pfizer Press Releases, 2022-Jan-18. ↩
4: S Ullrich, et al., “Main protease mutants of SARS-CoV-2 variants remain susceptible to nirmatrelvir (PF-07321332)”, bioRχiv, posted 2022-Jan-04. DOI: 10.1101/2021.11.28.470226. ↩
5: D Bojkova, et al., “Reduced interferon antagonism but similar drug sensitivity in Omicron variant compared to Delta variant SARS-CoV-2 isolates”, bioRχiv, posted 2022-Jan-04. ↩
6: L Vangeel, et al., “Remdesivir, Molnupiravir and Nirmatrelvir remain active against SARS-CoV-2 Omicron and other variants of concern”, bioRχiv, posted 2022-Jan-15. ↩
Gestae Commentaria
Comments for this post are closed pending repair of the comment system, but the Email/Twitter/Mastodon icons at page-top always work.